Short-read sequencing generates short DNA sequences (typically 50–300 base pairs). This technology is critical for disease research and supporting clinical diagnosis through whole-genome and exome sequencing. High-throughput short-read technology enables fast, cost-effective, and scalable analysis.
While short-read sequencing offers cost efficiency and scalability, its model accuracy may be limited when reconstructing complex genetic variations, exploring large structural variants, rare variants, or epigenetics. Since long-read sequencing can generate longer sequences, it has higher model accuracy in application scenarios such as oncology and other diseases.
By producing sequences of tens of thousands of base pairs, long reads can improve the model accuracy for identifying more polymorphisms (including insertions, deletions, and structural variants). Additionally, they enable more accurate genome assembly without the need for a reference genome.
For over a decade, scientists have used single-cell genomics to better understand biology and disease. By examining individual cells, researchers can observe various cellular states and their interactions. This helps them study gene expression, identify unique states, and pinpoint rare cell types associated with specific diseases.
Bulk RNA sequencing methods typically pool RNA from cells or tissues for analysis. Unlike baseline cell sequencing, which provides average expression levels across a sample, single-cell approaches enable granular, cell-level analysis. As a result, single-cell genomics can more precisely characterize variations between individual cells in control and disease samples.
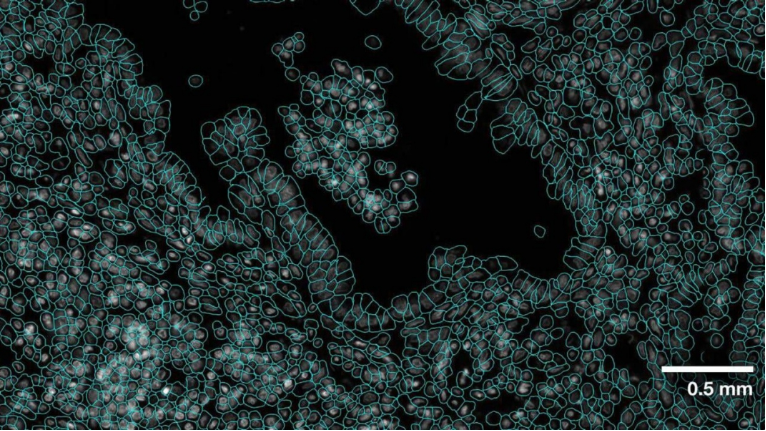
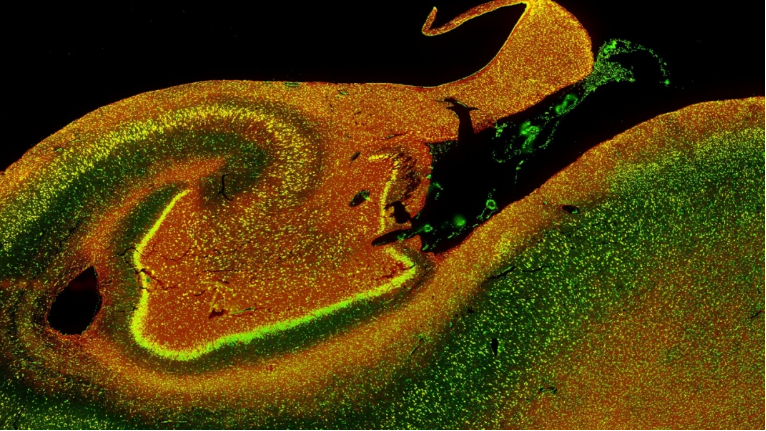
While single-cell technologies help researchers understand diseases by evaluating cells at the individual level, they lack the spatial context of the tissue surrounding these cells. With the advent of spatial transcriptomics, researchers can leverage all data—from relational data to imaging data—to better understand gene expression and cellular dynamics.
Relational data provides context for interactions between cells, enabling the overlay of imaging data with molecular data. The localization of cells and their interactions within the environment are critical for research, especially when studying rare cell types. Moreover, the context provided by spatial transcriptomics goes beyond local cellular interactions: it can show how diseases progress within the framework of the body’s systems. As a result, scientists can gain unprecedented spatial context about rare cell types and disease progression.